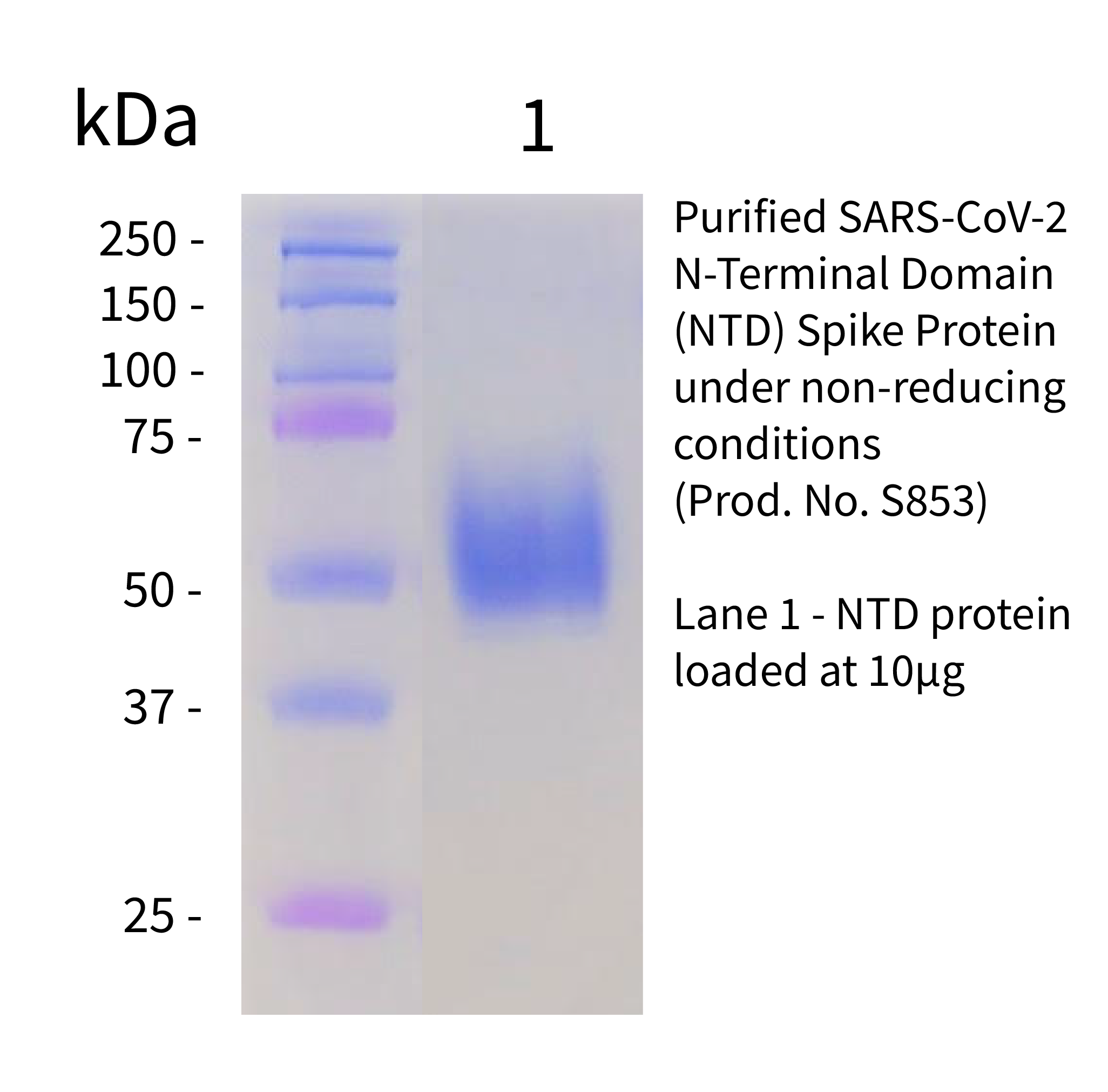
Recombinant SARS-CoV-2, NTD Protein
Data
- -
- -
BackgroundSevere acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative agent of coronavirus disease 2019 (COVID-19), is an enveloped, single-stranded, positive-sense RNA virus that belongs to the Coronaviridae family 1. The SARS-CoV-2 genome, which shares 79.6% identity with SARS-CoV, encodes four essential structural proteins: the spike (S), envelope (E), membrane (M), and nucleocapsid protein (N) 2. The S protein is a transmembrane, homotrimeric, class I fusion glycoprotein that mediates viral attachment, fusion, and entry into host cells 3. Each ~180 kDa monomer contains two functional subunits, S1 (~700 a.a) and S2 (~600 a.a), that mediate viral attachment and membrane fusion, respectively. S1 contains two major domains, the N-terminal (NTD) and C-terminal domains (CTD). In both SARS-CoV and SARS-CoV-2, the CTD contains the receptor-binding domain (RBD), which binds to the angiotensin-converting enzyme 2 (ACE2) receptor on host cells3-5. The NTD of SARS-CoV-2 does not bind to ACE26, and the function of NTD in SARS-CoV-2 infection is not well understood. In other CoVs, the NTD may promote attachment by binding to sugar moieties7 and might play a role in the conformational change of S2 required for membrane fusion8. While most neutralizing antibodies target the RBD domain and block receptor binding, potent neutralizing antibodies targeting NTD were isolated from convalescent COVID19 patients9, identifying the NTD as an attractive candidate for vaccines and therapeutics. In addition, the NTD is a promising antigen for diagnostic tests, as there is only 53.5% homology between the NTD of SARS-CoV-2 and SARS-CoV10. Protein DetailsFormat Purified No Carrier Protein Purity >95% by SDS Page Product Concentration 0.5 mg/ml Endotoxin Level <0.10 EU per 1 μg of the protein by the LAL method Protein Accession No. Amino Acid Sequence VNLTTRTQLPPAYTNSFTRGVYYPDKVFRSSVLHSTQDLFLPFFSNVTWFHAIHVSGTNGTKRFDNPVLPFNDGVYFASTEKSNIIRGWIFGTTLDSKTQSLLIVNNATNVVIKVCEFQFCNDPFLGVYYHKNNKSWMESEFRVYSSANNCTFEYVSQPFLMDLEGKQGNFKNLREFVFKNIDGYFKIYSKHTPINLVRDLPQGFSALEPLVDLPIGINITRFQTLLALHRSYLTPGDSSSGWTAGAAAYYVGYLQPRTFLLKYNENGTITDAVDCALDPLSETKCTLKS
State of Matter Sterile Liquid Predicted Molecular Mass The predicted molecular mass is ~36 kDa. Predicted Molecular Mass ~36 kDa Formulation This recombinant protein is aseptically packaged and formulated in 0.01 M phosphate buffered saline (PBS) pH 7.2 - 7.4, 150 mM NaCl with no carrier protein, potassium, calcium or preservatives added. Due to inherent biochemical properties of proteins, certain products may be prone to precipitation over time. Precipitation may be removed by aseptic centrifugation and/or filtration. Storage and Stability This recombinant protein may be stored as received at 2-8°C for up to one month. For longer term storage, aseptically aliquot in working volumes without diluting and store at -80°C. Avoid Repeated Freeze Thaw Cycles. Country of Origin USA Shipping Next Day NCBI Gene Bank Applications and Recommended Usage ? (Quality Tested by Leinco) ELISA References & Citations1. Zhou, P., Yang, X., Wang, X. et al. Nature 579, 270–273. 2020. 2. Wu, F., Zhao, S., Yu, B. et al. Nature 579, 265–269. 2020. 3. Wrapp D, Wang N, Corbett KS, et al. bioRxiv. 2020. 4. Walls AC, Park YJ, Tortorici MA, et al. Cell. 181(2):281-292.e6. 2020. 5. Li W, Zhang C, Sui J, et al. EMBO J. 24(8):1634-1643. 2020. 6. Wang Q, Zhang Y, Wu L, et al. Cell. 181(4):894-904.e9. 2020. 7. Park, Y., Walls, A.C., Wang, Z. et al. Nat Struct Mol Biol 26, 1151–1157. 2019. 8. Zhou, H., Chen, Y., Zhang, S. et al. Nat Commun 10, 3068. 2019. 9. Chi X, Yan R, Zhang J, et al. Science. eabc6952. 2020. 10. Ou, X., Liu, Y., Lei, X. et al. Nat Commun 11, 1620. 2020. Technical ProtocolsCertificate of AnalysisIMPORTANT Use lot specific datasheet for all technical information pertaining to this recombinant protein. |
Related Products
- -
- -
Prod No. | Description |
|---|---|
A314 | |
S540 | |
A313 | |
S538 | |
A315 | |
A316 | |
S536 | |
S537 | |
S539 | |
S541 | |
S542 | |
S543 | |
S544 |
 Products are for research use only. Not for use in diagnostic or therapeutic procedures.
Products are for research use only. Not for use in diagnostic or therapeutic procedures.